| Journal of Endocrinology and Metabolism, ISSN 1923-2861 print, 1923-287X online, Open Access |
| Article copyright, the authors; Journal compilation copyright, J Endocrinol Metab and Elmer Press Inc |
| Journal website https://jem.elmerpub.com |
Original Article
Volume 16, Number 1, February 2026, pages 29-37
Investigation of GHRL, RETN, ADIPOQ, NPY and MC4R Gene Polymorphisms in Obese and Non-Obese Patients With Diabetes
Saadet Busra Aksoyer Sezgina , Sermin Durakb
, Faruk Celikc, d
, Arezoo Gheybie
, Murat Diramalif
, Ramazan Cakmakg
, Mustafa Kerem Ozyavuze
, Ali Osman Gurolh
, Ilhan Yaylime
, Umit Zeybeke, i
aDepartment of Medical Biology and Genetics, Faculty of Medicine, Istanbul Yeni Yuzyil University, Istanbul 34010, Turkey
bDepartment of Medical Microbiology, Faculty of Medicine, Istanbul University Cerrahpasa, Istanbul 34320, Turkey
cDepartment of Medical Genetics, Faculty of Medicine, Alanya Alaaddin Keykubat University, Alanya, Antalya 07425, Turkey
dMolecular Medicine Research Center, Alanya Alaaddin Keykubat University, Alanya 07425, Antalya, Turkey
eDepartment of Molecular Medicine, Aziz Sancar Institute of Experimental Medicine, Istanbul University, Istanbul 34093, Turkey
fDepartment of Anatomy, Faculty of Medicine, Bolu Abant Izzet Baysal University, Bolu 14030, Turkey
gDepartment of Internal Decease, Istanbul University, Istanbul 34093, Turkey
hDepartment of Immunology, Aziz Sancar Institute of Experimental Medicine, Istanbul University, Istanbul 34093, Turkey
iCorresponding Author: Umit Zeybek, Department of Molecular Medicine, Aziz Sancar Institute of Experimental Medicine, Istanbul University, Istanbul 34093, Turkey
Manuscript submitted September 22, 2025, accepted October 27, 2026, published online February 25, 2026
Short title: GHRL, RETN, ADIPOQ, NPY, and MC4R Variants in DM
doi: https://doi.org/10.14740/jem1566
| Abstract | ▴Top |
Background: Obesity is a major public health issue caused by an abnormal increase in adipose tissue and is linked to increased insulin resistance and type 2 diabetes. Diabetes is a metabolic condition characterized by high glucose levels that is caused by insulin deficiency and alterations in insulin action mechanisms. Many of the genes associated with obesity are expressed in the central nervous system. Ghrelin, the endogenous ligand of the growth hormone secretagogue receptor, has a single-nucleotide polymorphism (SNP), rs4684677, which is associated with insulin resistance and metabolic syndrome. Adipose tissue secretes hormones known as adipokines, which include resistin and adiponectin, encoded by the ADIPOQ gene. SNPs in the RETN gene (rs1862513) have been linked to obesity and diabetes. Adiponectin is a cytokine generated from adipose tissue, and its polymorphism (rs2241766) increases the incidence of type 2 diabetes. Ghrelin binds to its receptor and promotes the expression of neuropeptide Y (NPY), thereby increasing food intake. NPY (rs16147) polymorphism has been linked to diabetes and obesity. The melanocortin 4 receptor (MC4R) is a regulator of energy balance and appetite. MC4R (rs17782313) polymorphism was associated with an increased risk of obesity.
Methods: Our study included 99 obese with diabetes and 99 non-obese patients with diabetes. RETN (rs1862513), GHRL (rs4684677), MC4R (rs17782313), NPY (rs16147), and ADIPOQ (rs2241766) genes were genotyped using real-time polymerase chain reaction.
Results: As a result, the RETN, MC4R, and NPY polymorphisms showed statistically significant differences in genotype/allele distributions between groups.
Conclusions: In conclusion, by representing data from the Turkish population for the first time, this study offers fresh insights and provides a unique perspective on the impact of the indicated genes on obesity, thereby contributing to the understanding of a hitherto unstudied ethnic group.
Keywords: Diabetes; ADIPOQ; NPY; MC4R; Resistin; Ghrelin
| Introduction | ▴Top |
Obesity is a global health issue because of its association with chronic diseases such as cardiovascular disease, type 2 diabetes mellitus, and cancer. Obesity also promotes the development of proinflammatory molecules that contribute to the pathophysiology of insulin resistance. Many of the genes associated with obesity are expressed in the central nervous system [1]. The leptin-melanocortin pathway is in charge of regulating hunger and energy metabolism. During periods of dietary restriction and low energy stores, the stomach secretes a large amount of ghrelin. Ghrelin, the endogenous ligand of the growth hormone secretagogue receptor, also plays an important role in body weight, insulin, and glucose metabolism. Single-nucleotide polymorphisms (SNPs) in the GHRL gene have been linked to obesity and diabetes in the literature, and these variants have been demonstrated to vary between populations [2–4].
Adipose tissue, an endocrine organ, secretes hormones known as adipokines, which include leptin, resistin, and adiponectin. In animals, the resistin protein is important in controlling glucose metabolism. SNPs in the RETN gene have been linked to metabolic, inflammatory, and autoimmune illnesses, as well as the development of insulin resistance [5], and some SNP variations have been linked to ethnicity [6, 7].
Adiponectin (ADIPOQ), an adipokine produced by adipocytes, has received substantial research for its anti-diabetic, anti-inflammatory, and anti-obesity properties [8–10]. Reduced blood adiponectin levels have been linked to decreased anti-inflammatory and enhanced pro-inflammatory actions, which contribute to the advancement of obesity-related disorders [11]. According to a study published in the literature, the ADIPOQ rs1501299 (276 G/T) polymorphism could be a candidate gene regulating obesity risk [12]. Ghrelin binds to the receptor and increases the expression of neuropeptide Y (NPY) and Agouti-related protein (AgRP), thus increasing food intake. Different SNPs in the NPY gene have been linked to obesity, glucose, and lipid metabolism from adolescence through young adulthood in the literature [13]. AgRP has been shown to operate as an antagonist to the melanocortin 4 receptor (MC4R), enhance appetite, and decrease energy expenditure [14–16]. MC4R is a major regulator of energy balance and appetite, and MC4R gene mutations are the most common monogenic cause of obesity in obese people, with a prevalence of 1.7–3.0% [17]. MC4R gene variations (p.N274S, p.S136F, p.V166I, p.R165W, p.I291SfsX10, p.M200del, p.S188L) are relatively prevalent in the Turkish population with childhood obesity, and screening for these variants is critical in children undergoing treatment for childhood obesity [18]. Another study in the literature found that the mutant MC4R rs17782313 was strongly related to the risk of obesity [19]. Furthermore, some experimental investigations have revealed that the effect of MC4R rs17782313 on obesity is ethnically dependent [20–23].
An examination of the literature revealed that no studies on the RETN, MC4R, NPY, and ADIPOQ genes in the Turkish population had been found. This study aimed to investigate the association of the RETN, GHRL, ADIPOQ, NPY, and MC4R polymorphisms with obesity and diabetes in a Turkish population for the first time, thereby filling a gap in the existing literature.
| Materials and Methods | ▴Top |
Ethical compliance
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Istanbul University Clinical Research Ethics Committee (Decree No. 13.05.2016/09).
Study design
Our study included 99 diabetic obese people and 99 diabetic non-obese people. The research was carried out at Istanbul University’s Aziz Sancar Experimental Medicine Research Institute, Department of Molecular Medicine. The first group included those who were diagnosed with diabetes using the Turkish Diabetes Association’s criteria and also met the criterion for obesity. They were chosen from the Department of Internal Medicine, Division of Endocrinology and Metabolism, Istanbul University Faculty of Medicine Hospital. The second group, on the other hand, was made up of diabetics who did not have obesity. Individuals with a history of chronic conditions such as cancer were excluded from the study. After describing the goal of the study, all participants signed informed permission forms. All of the participants were of Turkish descent. Individuals’ blood samples of 5 mL were collected, and the serums were maintained at –80 °C until the day of analysis after being centrifuged at 3,000 g for 15 min. Urea, creatinine, alanine aminotransferase (ALT), aspartate aminotransferase (AST), total cholesterol (TC), triglyceride (TG), high-density lipoprotein (HDL), low-density lipoprotein (LDL), glucose, and hemoglobin A1c (HbA1c) were all measured in the samples.
Genomic DNA isolation
Genomic DNA was extracted from 5 mL peripheral blood samples using a robotic system (RINATM M14; Bioeksen/Turkey). This machine separated DNA using magnetic beads, followed by chemical, physical, and enzymatic digestion, and then applied heat treatment with a subsequent washing step.
Real-time polymerase chain reaction (PCR)
RETN rs1862513, GHRL rs4684677, MC4R rs17782313, NPY rs16147, and ADIPOQ rs2241766 genes were genotyped using real-time PCR. Two forward and one reverse primer were designed for each SNP. Table 1 lists the primers used to amplify the relevant SNP. All reactions were carried out using a BioRad CFX96 instrument (Bio-Rad, Hercules, CA) and analyzed with the CFX Maestro Manager Software (Bio-Rad, Hercules, CA). The genotype distribution and allele frequencies of four gene polymorphisms in diabetic obese and non-obese diabetic groups were studied. The qPCR reactions contained 3 µL of oligo primer with Sybr Green, 5 µL of qPCR Master Mix (Bioeksen/Turkey), and 2 µL of template DNA. The total reaction volume was set for 10 µL for each primer. The qPCR protocol was followed: 3 min at 95 °C, and then by 40 cycles at 95 °C for 10 s, 60 °C for 30 s.
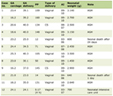 Click to view | Table 1. Primer Sequences for PCR Amplification of RETN rs1862513, GHRL rs4684677, MC4R rs17782313, NPY rs16147 and ADIPOQ rs2241766 Genes |
Statistical analysis
We used post-amplification fluorescence melting curve analysis to determine the allelic types of the SNP. Cq values obtained from standard curves and PCR were determined using CFX Manager Software version 3.1 (Bio-Rad Laboratories). SPSS 21.0 software (SPSS Inc., Chicago, IL, USA) was used to compare the demographic data of the groups. Statistical significance was defined as a P value less than 0.05.
The conformity of continuous data (Table 2) to normal distribution was assessed using the Shapiro-Wilk test. Data conforming to a normal distribution were presented as mean ± standard error (SE). The distribution of variances was examined using the Levene test. Differences between groups were assessed using the Student’s t-test or the Welch test. Categorical data (Table 3) were presented as frequencies and percentages. Differences between groups were assessed using the Chi-square test.
 Click to view | Table 2. Demographic and Clinical Data of Obese and Non-Obese Diabetic Groups |
 Click to view | Table 3. Genotype and Allele Frequencies of RETN rs1862513, GHRL rs4684677, MC4R rs17782313, NPY rs16147 and ADIPOQ rs2241766 Polymorphisms in the Study Groups |
| Results | ▴Top |
Results obtained from biochemical parameters
Our study is a case-control study with diabetic obese (n = 99, 50%) and non-obese diabetic (n = 99, 50%) patients. When we compared the groups in terms of biochemical and anthropological parameters, we discovered that the diabetic obese group had significantly higher age, urea, ALT, body mass index (BMI) and TG parameters (P < 0.05) (Table 2), while the non-obese diabetic group had significantly higher HDL parameters (P < 0.05) (Fig. 1). Other metrics showed no significant difference between groups (P > 0.05) (Table 2).
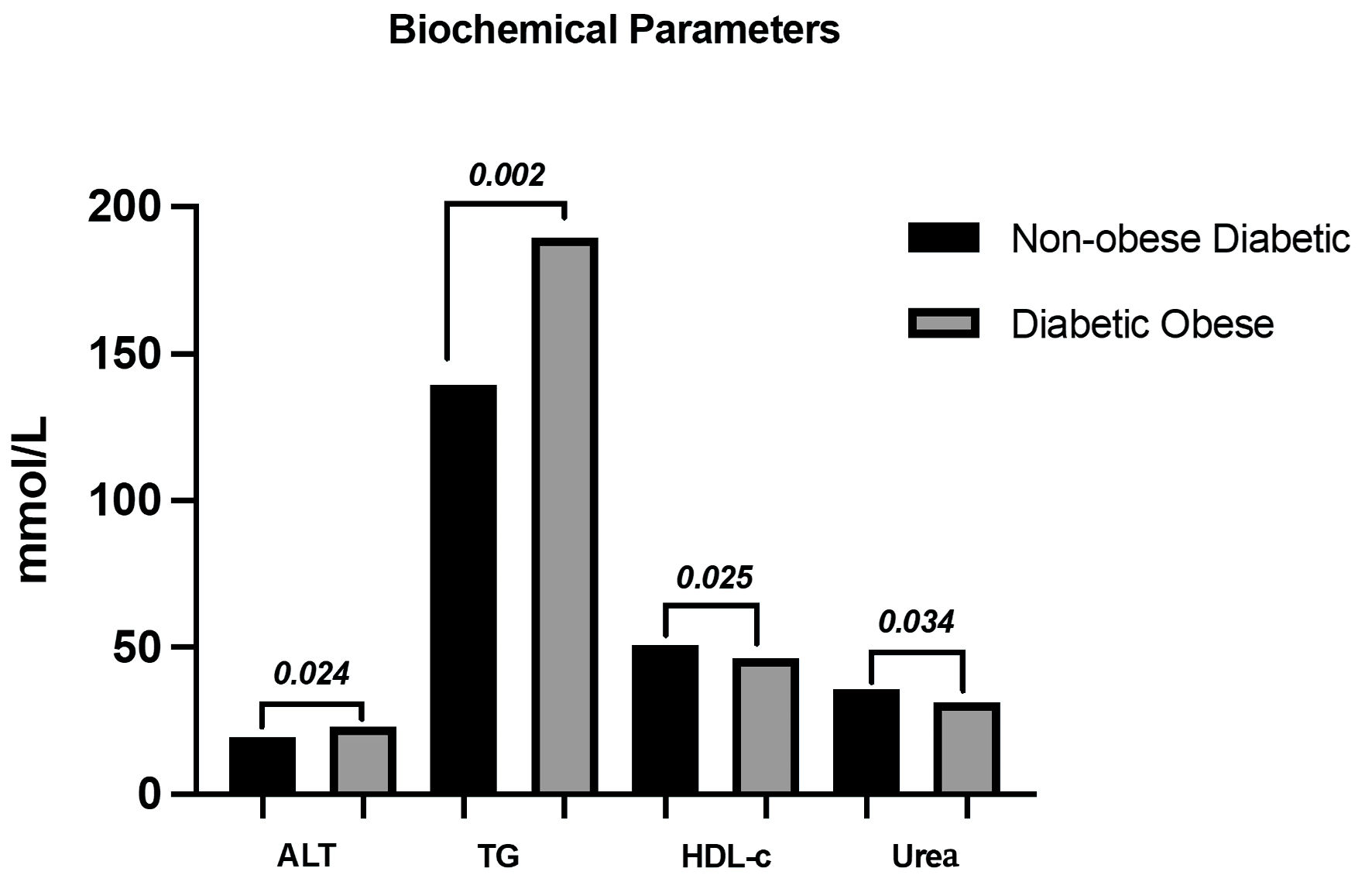 Click for large image | Figure 1. Biochemical parameters measured in the study groups. ALT: alanine aminotransferase; TG: triglyceride; HDL-C: high-density lipoprotein cholesterol. |
Results obtained from genotype and allele frequencies
The genotype and allele distributions of the RETN, MC4R, NPY, GHRL, and ADIPOQ polymorphisms were compared between groups in our study. The genotype distribution of RETN polymorphism was shown to be statistically significant between groups (P = 0.005). The diabetic obese group had a considerably greater C allele of the RETN polymorphism compared to the non-obese group (69.7% vs. 60.1%; P = 0.002). Furthermore, diabetic patients with the C allele of the RETN polymorphism had a nearly fourfold increased risk of obesity (odds ratio (OR) = 3.98; 95% confidence interval (CI) = 1.619–9.773).
The genotype distribution of the MC4R polymorphism was likewise shown to be statistically significant between groups (P = 0.008). The diabetic obese group had a higher and statistically significant C allele of the MC4R polymorphism (51% vs. 47%; P = 0.007). Diabetes patients with the C allele of the MC4R polymorphism had a nearly threefold increased incidence of obesity (OR = 3.33; 95% CI = 1.337–8.282).
The genotype distribution of the NPY polymorphism was statistically significant between groups (P < 0.0001). The diabetic obese group had a substantially greater T allele of the NPY polymorphism (62.1% vs. 45%; P < 0.0001), whereas the non-obese diabetic group had a higher and significantly different C allele (55% vs. 37.9%; P = 0.006). Furthermore, diabetic patients with the T allele of the NPY polymorphism had a 13-fold greater risk of obesity (OR = 13.058; 95% CI = 2.970–57.403). There was no significant difference in genotype and allele distribution between the groups for the GHRL and ADIPOQ polymorphisms (P > 0.05) (Table 3).
Biochemical parameters and findings from genotypes
Individuals having the C allele of the RETN polymorphism showed higher TC levels in the non-obese diabetic group (mean ± SE = 207.47 ± 6.5, P = 0.018). Individuals with the GHRL polymorphism’s A allele had statistically substantially higher TC levels (mean SE = 201.79 ± 5.5, P = 0.033), while those with the C allele had a higher age parameter (mean SE = 51.34 ± 1.5, P = 0.025). Individuals with the TT genotype of the MC4R polymorphism showed a substantial rise in TG levels in the non-obese diabetic group (mean SE = 179 ± 29.2, P = 0.017). Individuals with the G allele of the ADIPOQ polymorphism had significantly lower HbA1c levels (mean SD = 7.41 ± 0.36, P = 0.044).
There was no significant relationship between genotype and allele frequencies of the NPY polymorphism and metabolic markers in the non-obese diabetic group (Fig. 2).
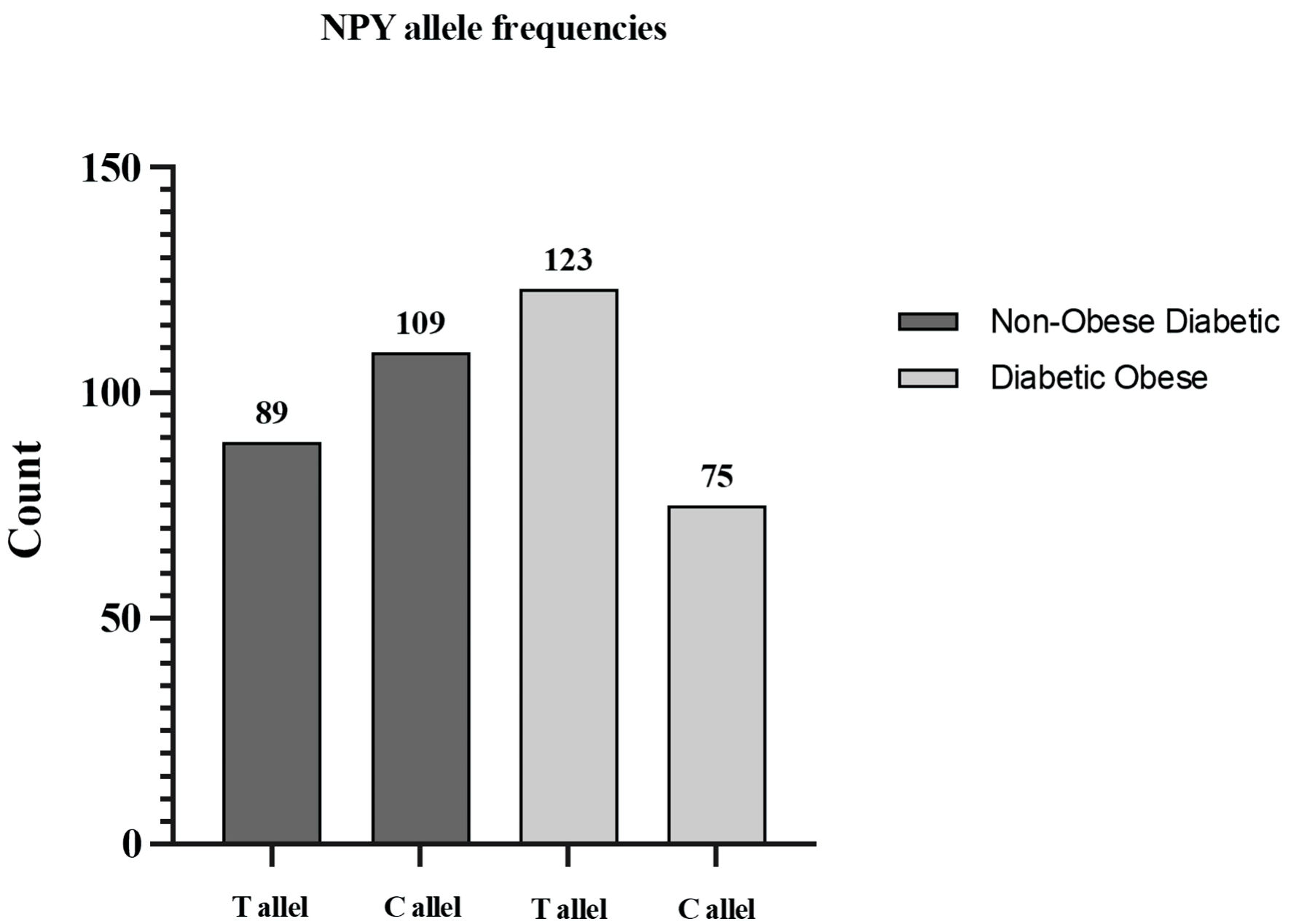 Click for large image | Figure 2. NPY allele distribution in study groups. |
Individuals with the T allele of the MC4R polymorphism had substantially higher levels of glucose (mean SE = 185.75 ± 7.2; P < 0.001) and HbA1c (mean SE = 8.4 ± 0.2; P = 0.022) in the diabetes obese group. Diabetic obese people with the C allele of the NPY polymorphism had a significant rise in BMI (mean SE = 37.1 ± 0.5; P < 0.001). Furthermore, diabetic obese people with the T allele of the ADIPOQ polymorphism had a significant rise in BMI (mean SE = 36.4 ± 0.4; P < 0.001). There was no statistically significant connection between genotypes and allelic distributions of RETN, GHRL, and ADIPOQ polymorphisms and biochemical markers in the diabetic obese group (P > 0.005).
| Discussion | ▴Top |
Obesity is a major public health issue that is becoming more prevalent around the world. It is caused by an abnormal increase in adipose tissue and is linked to increased insulin resistance, type 2 diabetes, cardiovascular disease, and non-alcoholic fatty liver disease. Diabetes, on the other hand, is a metabolic condition characterized by high blood glucose levels that are caused by insulin deficiency and alterations in insulin action mechanisms [24].
Ghrelin, a 28-amino acid gastrointestinal peptide, and its receptor are found in many peripheral organs, intestines, pancreatic islets, and brain regions. Gln90Leu rs4684677 is one of the SNPs associated with insulin resistance and metabolic syndrome. A study in the Spanish population discovered a link between the Ghrelin rs4684677 polymorphism and metabolic syndrome. However, no correlation was identified between genotype haplotypes and BMI [25].
The GHRL rs4684677 polymorphism has been reported to affect insulin sensitivity and eating behavior in different population studies in the literature [26, 27], but no association with gestational diabetes or type 2 diabetes [28]. Similarly, no significant differences were observed between the genotype and allele distributions of GHRL rs4684688 and biochemical parameters in the study groups and the diabetic obese group (P > 0.05). These findings in our study with diabetic obese and non-obese diabetic individuals are consistent with the results of relevant publications in literature.
Resistin is an adipokine produced by adipose tissue and is a member of the cysteine-rich secretory protein family. Obesity, insulin resistance, and type 2 diabetes have all been linked to SNPs in the RETN gene and elevated resistin concentrations in serum [29, 30]. The GG genotype of rs1862513 was indicated to be an indication of lower homeostasis model assessment of insulin resistance (HOMA-IR), insulin, and LDL cholesterol in a study conducted by De Luis et al in non-diabetic obese patients [30].
However, in different studies conducted in different populations, the rs182513 polymorphism was found to have no effect on metabolic syndrome susceptibility in individuals with obesity and diabetes, and no significant relationship was found between the allelic frequencies of rs1862513 and the genotypic distributions with metabolic syndrome [31, 32].
Contrary to the literature, the genotype distribution of the RETN rs1862513 SNP was shown to be significant in our investigation, and it was reported that carrying the C allele enhanced the risk of obesity in diabetic patients by four times. These findings show that people with the C allele are more likely to develop metabolic syndrome. Individuals having the C allele had significantly higher TC levels (P < 0.05) in the non-obese diabetic group; however, no significance was observed when assessing genotype and allele distributions in terms of biochemical markers in the diabetic obese group. These data show that having high TC among non-obese diabetics with the C allele may raise the risk of cardiovascular disease; however, further studies are needed to confirm this finding.
Adiponectin is a cytokine generated from adipose tissue that influences diabetes. ADIPOQ rs2241766 polymorphism increases the incidence of type 2 diabetes [33] and is highly related to diabetes [34–39]. Studies in diverse populations have also found that the ADIPOQ rs2241766 polymorphism is linked to the risk of obesity [40] and the development of type 2 diabetes [10, 38, 41, 42].
In contrast to these findings, there are studies in the literature that show no link between the ADIPOQ rs2241766 polymorphism and type 2 diabetes [43] or obesity risk [44].
In our investigation, similar to the findings of Lu et al [44], no significant differences in the genotype and allele distributions of the ADIPOQ rs2241766 polymorphism were detected in diabetic obese and non-obese diabetic groups. In terms of biochemical parameters, however, individuals carrying the G allele in the non-obese diabetic group had decreased and significant HbA1c levels, whereas there was no significant difference in terms of genotype and allele in the diabetic obese group.
NPY is a 36-amino acid peptide neurotransmitter produced by chromaffin cells and noradrenergic neurons. The NPY rs16147 SNP has been linked to type 2 diabetes and obesity risk [45], and it has been proposed that this is due to enhanced NPY signaling caused by elevated NPY expression in the hypothalamus [46]. BMI, fat mass, waist circumference, leptin, and insulin levels were considerably lower in males bearing the A allele in a study on NPY rs16147 polymorphism in obese patients [47]. The rs16139 SNP has been found to be strongly associated with obesity measures such as BMI and waist circumference, and the T allele is associated with an increased risk of obesity [48, 49]. This points to a consistent link between rs16147 and obesity risk in Caucasians and Asians.
Similarly, the NPY rs16147 genotype distribution was shown to be significant in our study, and the C allele is high and significant in the non-obese diabetes group, whereas the T allele is high and significant in the diabetic obese group. Furthermore, those with the T allele had a 13-fold increased risk of obesity. The correlation of the NPY SNP with obesity is extremely consistent in the populations analyzed in the literature, and our population’s homogeneity also suggests a genetically connected link. In terms of biochemical markers, no significance was found in the genotype and allele distributions in the non-obese diabetic group; however, it was established that BMI was greater and significant in persons carrying the C allele in the diabetic obese group. Although the mechanism behind the link between the NPY variation and increased BMI is thought to be increased food consumption, there is limited evidence from epidemiological research to support this idea. More research is needed to understand the molecular mechanisms behind the observed relationships.
MC4R mutations are the most prevalent monogenic cause of obesity, accounting for 1.7–3.0% of obese people [17]. The most commonly researched MC4R gene variant, rs17782313, has been linked to obesity [50, 51], and has substantial associations with calorie consumption [52], increased snacking, and hunger [53]. Several studies in diverse populations have found that the MC4R rs17782313 polymorphism is related to an increased risk of obesity and diabetes [19, 54, 55, 56]. Another study in overweight and obese adults discovered that different MC4R rs17782313 genotypes were related to energy consumption and the C allele [57]. Similarly, we discovered that the genotype distribution and C allele of MC4R rs17782313 were considerably greater in diabetic obese individuals in our study. As a result, diabetes patients having the C allele have a threefold increased risk of obesity. When we looked at the biochemical markers, we found that patients with the TT genotype had considerably higher TG levels in the non-obese diabetic group, while individuals with the T allele had higher and significant levels of glucose and HbA1c in the diabetic obese group. This suggests that non-obese diabetics are more likely to acquire cardiovascular disease. Our work is the first to look at the combined effects of RETN, GHRL, MC4R, NPY, and ADIPOQ polymorphisms in diabetic patients with and without obesity. The findings of this study add significantly to the existing body of knowledge regarding the phenotypic impact of these disorders. Furthermore, to our knowledge, this is the first scientific study conducted in Turkey to examine all of these genes within a different ethnic group.
Acknowledgments
The authors would like to thank TUBITAK (Project No: 116E827) for its financial support.
Financial Disclosure
This study was supported by the Research Fund of the Scientific and Technological Research Council of Turkey (TUBITAK 1003, Project No: 116E827).
Conflict of Interest
The authors declare no competing interests.
Informed Consent
Informed consent was obtained from all individual participants included in the study. Patients signed informed consent regarding publishing their data.
Author Contributions
All authors contributed to the study conception and design. Study concept and design: Saadet Busra Aksoyer Sezgin and Umit Zeybek; acquisition of data: Saadet Busra Aksoyer Sezgin, Sermin Durak, Ramazan Cakmak, Mustafa Kerem Ozyavuz; analysis and interpretation of data: Saadet Busra Aksoyer Sezgin, Umit Zeybek, Sermin Durak, Faruk Celik and Arezoo Gheybi; drafting of the manuscript: Saadet Busra Aksoyer Sezgin, Sermin Durak and Faruk Celik; critical revision of the manuscript: Umit Zeybek, Ali Osman Gurol and Ilhan Yaylim; statistical analysis: Murat Diramali; study supervision: Umit Zeybek.
Data Availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
| References | ▴Top |
- Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, Lango Allen H, et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet. 2010;42(11):937-948.
doi pubmed - Becer E, Ergoren MC. Dual effect of the GHRL gene variant in the molecular pathogenesis of obesity. Balkan J Med Genet. 2021;24(1):27-34.
doi pubmed - Giha HA, Joatar FE, AlDehaini DMB, Malalla ZHA, Ali ME, Al Qarni AA. Association of obesity in T2DM with differential polymorphism of ghrelin, growth hormone secretagogue receptor-1 and telomeres maintenance genes. Horm Mol Biol Clin Investig. 2022;43(3):297-306.
doi pubmed - Imaizumi T, Ando M, Nakatochi M, Yasuda Y, Honda H, Kuwatsuka Y, Kato S, et al. Effect of dietary energy and polymorphisms in BRAP and GHRL on obesity and metabolic traits. Obes Res Clin Pract. 2018;12(Suppl 2):39-48.
doi pubmed - Tripathi D, Kant S, Pandey S, Ehtesham NZ. Resistin in metabolism, inflammation, and disease. FEBS J. 2020;287(15):3141-3149.
doi pubmed - El-Shal AS, Pasha HF, Rashad NM. Association of resistin gene polymorphisms with insulin resistance in Egyptian obese patients. Gene. 2013;515(1):233-238.
doi pubmed - Park HK, Ahima RS. Resistin in rodents and humans. Diabetes Metab J. 2013;37(6):404-414.
doi pubmed - Kim S, Lee Y, Kim JW, Son YJ, Ma MJ, Um JH, Kim ND, et al. Discovery of a novel potent peptide agonist to adiponectin receptor 1. PLoS One. 2018;13(6):e0199256.
doi pubmed - Pawlik A, Teler J, Maciejewska A, Sawczuk M, Safranow K, Dziedziejko V. Adiponectin and leptin gene polymorphisms in women with gestational diabetes mellitus. J Assist Reprod Genet. 2017;34(4):511-516.
doi pubmed - Biswas D, Vettriselvi V, Choudhury J, Jothimalar R. Adiponectin gene polymorphism and its association with type 2 diabetes mellitus. Indian J Clin Biochem. 2011;26(2):172-177.
doi pubmed - Komiyama M, Wada H, Yamakage H, Satoh-Asahara N, Sunagawa Y, Morimoto T, Ozaki Y, et al. Analysis of changes on adiponectin levels and abdominal obesity after smoking cessation. PLoS One. 2018;13(8):e0201244.
doi pubmed - Elghazy AM, Elsaeid AM, Refaat M, Youssef MM. Biochemical studies of adiponectin gene polymorphism in patients with obesity in Egyptians. Arch Physiol Biochem. 2022;128(1):43-50.
doi pubmed - Katus U, Villa I, Ringmets I, Veidebaum T, Harro J. Neuropeptide Y gene variants in obesity, dietary intake, blood pressure, lipid and glucose metabolism: A longitudinal birth cohort study. Peptides. 2021;139:170524.
doi pubmed - Larder R, Lim CT, Coll AP. Genetic aspects of human obesity. Handb Clin Neurol. 2014;124:93-106.
doi pubmed - Sutton AK, Myers MG, Jr., Olson DP. The role of PVH circuits in leptin action and energy balance. Annu Rev Physiol. 2016;78:207-221.
doi pubmed - Vetter ML, Faulconbridge LF, Webb VL, Wadden TA. Behavioral and pharmacologic therapies for obesity. Nat Rev Endocrinol. 2010;6(10):578-588.
doi pubmed - Kohlsdorf K, Nunziata A, Funcke JB, Brandt S, von Schnurbein J, Vollbach H, Lennerz B, et al. Early childhood BMI trajectories in monogenic obesity due to leptin, leptin receptor, and melanocortin 4 receptor deficiency. Int J Obes (Lond). 2018;42(9):1602-1609.
doi pubmed - Aykut A, Ozen S, Goksen D, Ata A, Onay H, Atik T, Darcan S, et al. Melanocortin 4 receptor (MC4R) gene variants in children and adolescents having familial early-onset obesity: genetic and clinical characteristics. Eur J Pediatr. 2020;179(9):1445-1452.
doi pubmed - Yu K, Li L, Zhang L, Guo L, Wang C. Association between MC4R rs17782313 genotype and obesity: A meta-analysis. Gene. 2020;733:144372.
doi pubmed - Srivastava A, Mittal B, Prakash J, Srivastava P, Srivastava N. Analysis of MC4R rs17782313, POMC rs1042571, APOE-Hha1 and AGRP rs3412352 genetic variants with susceptibility to obesity risk in North Indians. Ann Hum Biol. 2016;43(3):285-288.
doi pubmed - Xi B, Chandak GR, Shen Y, Wang Q, Zhou D. Association between common polymorphism near the MC4R gene and obesity risk: a systematic review and meta-analysis. PLoS One. 2012;7(9):e45731.
doi pubmed - Hester JM, Wing MR, Li J, Palmer ND, Xu J, Hicks PJ, Roh BH, et al. Implication of European-derived adiposity loci in African Americans. Int J Obes (Lond). 2012;36(3):465-473.
doi pubmed - Grant SF, Bradfield JP, Zhang H, Wang K, Kim CE, Annaiah K, Santa E, et al. Investigation of the locus near MC4R with childhood obesity in Americans of European and African ancestry. Obesity (Silver Spring). 2009;17(7):1461-1465.
doi pubmed - Shah VN, Prattichizzo F, Ceriello A. Obesity and diabetes. Diabetes Technol Ther. 2023;25(S1):S217-S226.
doi pubmed - Mora M, Adam V, Palomera E, Blesa S, Diaz G, Buquet X, Serra-Prat M, et al. Ghrelin gene variants influence on metabolic syndrome components in aged Spanish population. PLoS One. 2015;10(9):e0136931.
doi pubmed - Nicklas TA, Baranowski T, Cullen KW, Berenson G. Eating patterns, dietary quality and obesity. J Am Coll Nutr. 2001;20(6):599-608.
doi pubmed - Gueorguiev M, Lecoeur C, Meyre D, Benzinou M, Mein CA, Hinney A, Vatin V, et al. Association studies on ghrelin and ghrelin receptor gene polymorphisms with obesity. Obesity (Silver Spring). 2009;17(4):745-754.
doi pubmed - Rocha RA, Frigeri HR, Santos-Weiss IC, Rea RR, Souza EM, Rego FG, Picheth G. Preproghrelin polymorphism Q90L (rs4684677) in gestational diabetes. Arq Bras Endocrinol Metabol. 2014;58(1):83-84.
doi pubmed - Meier U, Gressner AM. Endocrine regulation of energy metabolism: review of pathobiochemical and clinical chemical aspects of leptin, ghrelin, adiponectin, and resistin. Clin Chem. 2004;50(9):1511-1525.
doi pubmed - de Luis DA, Izaola O, Primo D, Aller R. Effect of the rs1862513 variant of resistin gene on insulin resistance and resistin levels after two hypocaloric diets with different fat distribution in subjects with obesity. Eur Rev Med Pharmacol Sci. 2018;22(12):3865-3872.
doi pubmed - Suriyaprom K, Tungtrongchitr R, Namjuntra P. Associations of resistin levels with resistin gene polymorphism and metabolic syndrome in Thais. J Med Biochem. 2015;34(2):170-178.
doi pubmed - Fu Y, Yu Y, Wu Y, You Y, Zhang Y, Kou C. Association between two resistin gene polymorphisms and metabolic syndrome in Jilin, Northeast China: A Case-Control Study. Dis Markers. 2017;2017:1638769.
doi pubmed - Hamidi Y, Saki S, Afraz ES, Pashapour S. A Meta-analysis of ADIPOQ rs2241766 polymorphism association with type 2 diabetes. J Diabetes Metab Disord. 2022;21(2):1895-1901.
doi pubmed - Foucan L, Maimaitiming S, Larifla L, Hedreville S, Deloumeaux J, Joannes MO, Blanchet-Deverly A, et al. Adiponectin gene variants, adiponectin isoforms and cardiometabolic risk in type 2 diabetic patients. J Diabetes Investig. 2014;5(2):192-198.
doi pubmed - Nezamzadeh F, Esmailkhani A, Edalati E,Hosseini SS, Ghasemi A, Bialvaei AZ, Taheri K. Link between single nucleotide polymor- phism of rs266729 and rs2241766 in the ADIPOQ gene and gestational diabetes in an Iranian population. Gene Reports. 2019;14:72-75.
- Tu Y, Yu Q, Fan G, Yang P, Lai Q, Yang F, Zhang S, et al. Assessment of type 2 diabetes risk conferred by SNPs rs2241766 and rs1501299 in the ADIPOQ gene, a case/control study combined with meta-analyses. Mol Cell Endocrinol. 2014;396(1-2):1-9.
doi pubmed - Howlader M, Sultana MI, Akter F, Hossain MM. Adiponectin gene polymorphisms associated with diabetes mellitus: A descriptive review. Heliyon. 2021;7(8):e07851.
doi pubmed - Hussain MK, Deli FA, Algenabi AHA, Abdul-Rudha KH. Adiponectin gene polymorphisms as a predictor for development of type 2 diabetes mellitus in Iraqi population. Gene. 2018;662:118-122.
doi pubmed - Mtiraoui N, Ezzidi I, Turki A, Chaieb A, Mahjoub T, Almawi WY. Single-nucleotide polymorphisms and haplotypes in the adiponectin gene contribute to the genetic risk for type 2 diabetes in Tunisian Arabs. Diabetes Res Clin Pract. 2012;97(2):290-297.
doi pubmed - Wu J, Liu Z, Meng K, Zhang L. Association of adiponectin gene (ADIPOQ) rs2241766 polymorphism with obesity in adults: a meta-analysis. PLoS One. 2014;9(4):e95270.
doi pubmed - Hara K, Boutin P, Mori Y, Tobe K, Dina C, Yasuda K, Yamauchi T, et al. Genetic variation in the gene encoding adiponectin is associated with an increased risk of type 2 diabetes in the Japanese population. Diabetes. 2002;51(2):536-540.
doi pubmed - Toy W, Liu J, Cheng A, Tan CSH, Lau DP, Wong MDS, Subramaniam T, et al. Adiponectin gene polymorphisms and type 2 diabetes among Singaporean Chinese adults. J Diabetes Metabol. 2011;2(8).
doi - Potapov VA, Chistiakov DA, Dubinina A, Shamkhalova MS, Shestakova MV, Nosikov VV. Adiponectin and adiponectin receptor gene variants in relation to type 2 diabetes and insulin resistance-related phenotypes. Rev Diabet Stud. 2008;5(1):28-37.
doi pubmed - Lu JF, Zhou Y, Huang GH, Jiang HX, Hu BL, Qin SY. Association of ADIPOQ polymorphisms with obesity risk: a meta-analysis. Hum Immunol. 2014;75(10):1062-1068.
doi pubmed - Patel R, Dwivedi M, Mansuri MS, Ansarullah, Laddha NC, Thakker A, Ramachandran AV, et al. Association of Neuropeptide-Y (NPY) and Interleukin-1beta (IL1B), genotype-phenotype correlation and plasma lipids with type-II diabetes. PLoS One. 2016;11(10):e0164437.
doi pubmed - Sitticharoon C, Chatree S, Churintaraphan M. Expressions of neuropeptide Y and Y1 receptor in subcutaneous and visceral fat tissues in normal weight and obese humans and their correlations with clinical parameters and peripheral metabolic factors. Regul Pept. 2013;185:65-72.
doi pubmed - de Luis DA, Izaola O, de la Fuente B, Primo D, Aller R. Association of neuropeptide Y gene rs16147 polymorphism with cardiovascular risk factors, adipokines, and metabolic syndrome in patients with obesity. J Nutrigenet Nutrigenomics. 2016;9(5-6):213-221.
doi pubmed - Yeung EH, Zhang C, Chen J, Bowers K, Hu FB, Kang G, Qi L. Polymorphisms in the neuropeptide Y gene and the risk of obesity: findings from two prospective cohorts. J Clin Endocrinol Metab. 2011;96(12):E2055-2062.
doi pubmed - Zain SM, Mohamed Z, Jalaludin MY, Fauzi F, Hamidi A, Zaharan NL. Comprehensive evaluation of the neuropeptide-Y gene variants in the risk of obesity: a case-control study and meta-analysis. Pharmacogenet Genomics. 2015;25(10):501-510.
doi pubmed - Beckers S, Zegers D, de Freitas F, Mertens IL, Van Gaal LF, Van Hul W. Association study of MC4R with complex obesity and replication of the rs17782313 association signal. Mol Genet Metab. 2011;103(1):71-75.
doi pubmed - Corella D, Ortega-Azorin C, Sorli JV, Covas MI, Carrasco P, Salas-Salvado J, Martinez-Gonzalez MA, et al. Statistical and biological gene-lifestyle interactions of MC4R and FTO with diet and physical activity on obesity: new effects on alcohol consumption. PLoS One. 2012;7(12):e52344.
doi pubmed - Khalilitehrani A, Ghorbani M, Hosseini S, Pishva H. Association between physical activity and body mass index in different melanocortin receptor 4 (MC4R) genotypes. J Public Health Res. 2015;13(3):53-64.
- Stutzmann F, Cauchi S, Durand E, Calvacanti-Proenca C, Pigeyre M, Hartikainen AL, Sovio U, et al. Common genetic variation near MC4R is associated with eating behaviour patterns in European populations. Int J Obes (Lond). 2009;33(3):373-378.
doi pubmed - Renstrom F, Payne F, Nordstrom A, Brito EC, Rolandsson O, Hallmans G, Barroso I, et al. Replication and extension of genome-wide association study results for obesity in 4923 adults from northern Sweden. Hum Mol Genet. 2009;18(8):1489-1496.
doi pubmed - El Hajj Chehadeh S, Osman W, Nazar S, Jerman L, Alghafri A, Sajwani A, Alawlaqi M, et al. Implication of genetic variants in overweight and obesity susceptibility among the young Arab population of the United Arab Emirates. Gene. 2020;739:144509.
doi pubmed - Schreyer KF, Leucht S, Heres S, Steimer W. Genetic association of the rs17782313 polymorphism with antipsychotic-induced weight gain. Psychopharmacology (Berl). 2023;240(4):899-908.
doi pubmed - Rahati S, Qorbani M, Naghavi A, Pishva H. Association and interaction of the MC4R rs17782313 polymorphism with plasma ghrelin, GLP-1, cortisol, food intake and eating behaviors in overweight/obese Iranian adults. BMC Endocr Disord. 2022;22(1):234.
doi pubmed
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, including commercial use, provided the original work is properly cited.
Journal of Endocrinology and Metabolism is published by Elmer Press Inc.